No coding, no problem: Revolutionizing scientific workflows through workflow-orchestrating AI
2 Center for Systems Integration and Sustainability, Michigan State University, East Lansing, MI
HYW: https://orcid.org/0000-0002-2146-8257
LH: https://orcid.org/0000-0002-7422-0343
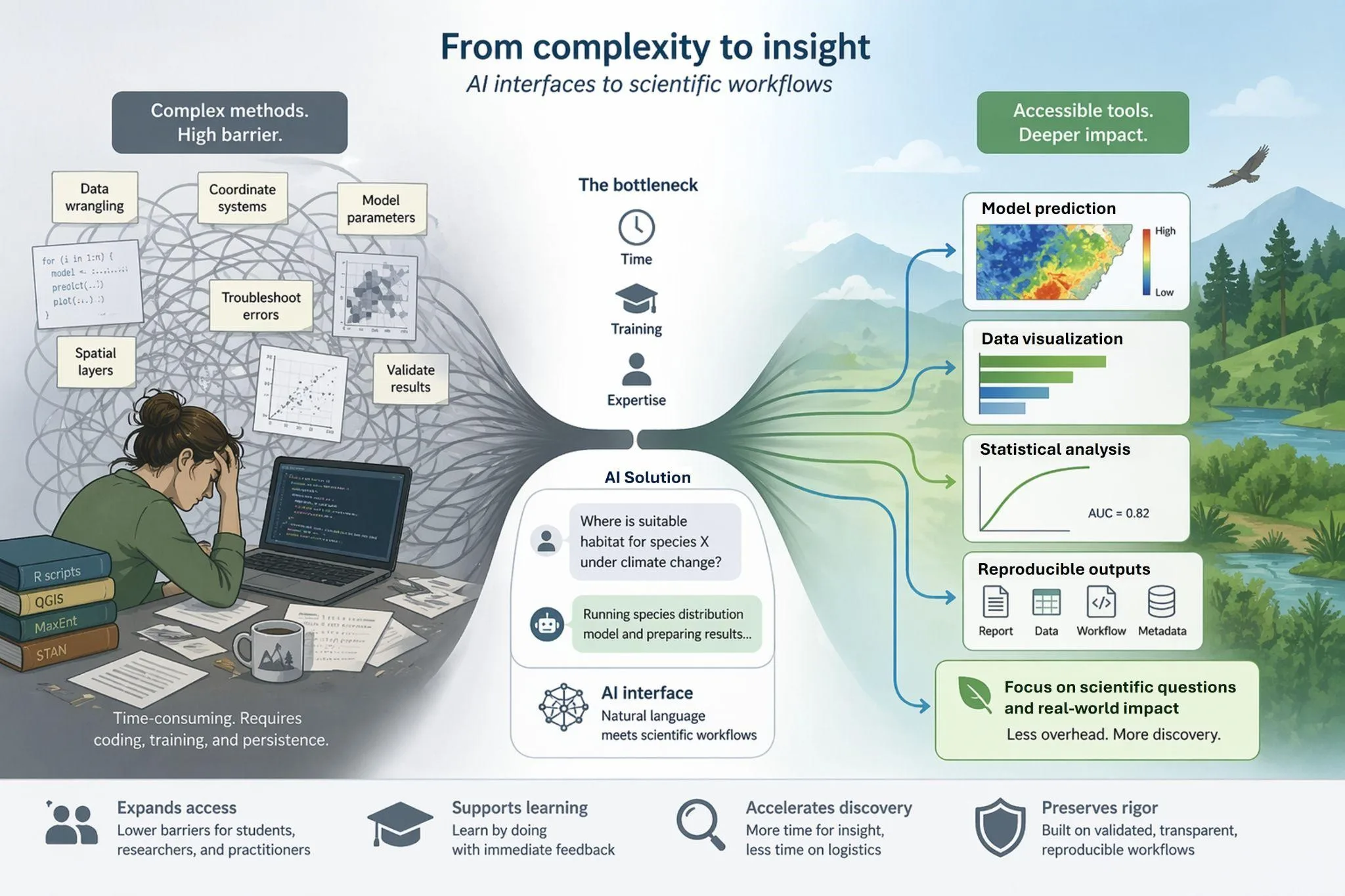
Abstract photo. Intelligent AI interfaces remove the need for coding by translating natural-language inputs into validated scientific workflows, reducing barriers in time, training, and expertise while enabling reproducible analyses and accelerating scientific insight.
Abstract
Scientists are confronting a widening gap between the complexity of environmental challenges and the capacity to efficiently respond. As global issues like climate change, biodiversity loss, and spatial planning become increasingly intricate, so do the methods and analytical tools required to address them. The time and training to master these methods have become a costly currency that many students and even seasoned researchers cannot afford. This results in a bottleneck that limits the number of challenges that can be addressed. We argue that emerging natural-language AI tools, powered by large language models, offer a transformative path forward when designed as interfaces to existing scientific workflows. By enabling researchers to interact with complex modeling frameworks through plain language, these systems can dramatically lower the execution overhead of advanced analyses while preserving transparency and reproducibility. This shift allows researchers to focus on ecological insight rather than coding logistics. We emphasize that AI is not a replacement for domain expertise or established methods, but can support systems that can expand access to complex analytical tools, support education, and accelerate discovery across ecology and other data-intensive disciplines in a time of urgent need.
Keywords: artificial intelligence, generative AI, interdisciplinary science, large language models, natural language, reproducibility
The expanding gap in research capacity
Scientists are entering a paradoxical era. On one hand, global challenges today are urgent, complex, and multiplying (Ripple et al., 2020). Climate change, biodiversity loss, habitat fragmentation, and emerging zoonotic diseases all demand fast, scalable insight from spatial and statistical models (Urban et al., 2016). On the other hand, the capacity of individual researchers and institutions to efficiently address these questions, whether it be time or resources, is trailing behind (Hampton et al., 2013, Soranno and Schimel, 2014). This is not because of apathy or a lack of ambition, but because the methodological demands of modern science are outpacing our ability to meet them.
Earth is complex, and problem solving often requires equally complex methods, which might require running complex simulations, fitting hierarchical models, or integrating spatial data with remote sensing. These approaches are now essential to credible, actionable science, but they require time, training, and interdisciplinary fluency that most research groups, especially students and early career scientists, cannot afford. The bottleneck is less the availability of ideas or methods, and more the time and expertise required to execute complex, fragmented workflows reliably.
More questions, more methods, less time
This mismatch is acute for early-career researchers, interdisciplinary teams, and conservation practitioners. As new modeling frameworks, packages, and software platforms emerge, the learning curve steepens, and would-be users can feel pulled in too many directions. Modern tools may be revolutionary, but each demands dozens of hours to learn, hundreds of lines of code to deploy, and substantial trial and error to adapt to a new dataset or question. Even seasoned researchers often spend more time debugging workflows than interpreting results.
For example, ecologists often use species distribution models (SDMs) to understand how a species might be distributed across a landscape. Conducting an SDM analysis demands fluency across a wide array of tools and concepts – statistical modeling (e.g., GLMs, MaxEnt), geospatial processing (e.g., GIS software, raster manipulation), data cleaning, performance evaluation (e.g., AUC, TSS), and computer programming in R or Python. These workflows are not only fragmented but are also becoming increasingly complex with the rise of machine learning and high-dimensional data. As a result, the barrier to entry is steep and often requires months or even years of training before a student or researcher can independently conduct meaningful analysis.
Meanwhile, global challenges are mounting faster than we can respond (Ripple et al., 2020). For ecologists, restoration planning, spatial prioritization, genomics and connectivity modeling, climate-driven range shifts, and landscape-scale conservation efforts require methodological agility that our current systems (i.e., institutional, educational) do not support within reasonable time frames (Moilanen et al., 2009, Sabo et al., 2024). As a result, critical questions go unanswered not due to a lack of data or insight, but because the tools remain out of reach. We must lower the barriers and streamline the workflows to use complex tools to meet the ecological and environmental challenges of the near future.
The rising potential of natural-language AI systems
Researchers are already using AI to support ecological analyses, for example through code comprehension, debugging, and drafting analysis plans. These applications can reduce friction for learners and interdisciplinary teams by making existing workflows more interpretable. At the same time, many AI advances in ecology have centered on building new machine-learning pipelines that require additional expertise, which can shift barriers rather than eliminate them.
Here we focus on a different opportunity, which is using AI to lower the execution overhead of established ecological workflows by making them accessible through governed natural-language interfaces rather than creating entirely new modeling methods.
Instead of forcing researchers to translate their questions into code, what if existing tools could understand plain language? What if asking “Compare the edge density of forest and grassland” or “Run a connectivity analysis” was enough to launch a reproducible, transparent ecological workflow?
Generative AI as a scientific aide and its limitations
Recent advances in large language models (LLMs) make natural-language interaction with computational systems possible. An LLM is a specialized AI model designed to process and generate text and forms the foundation of many natural-language AI systems. When these systems are used to generate text or code in response to user prompts, they are commonly described as Generative AI (GenAI; Feuerriegel et al. 2024).
GenAI systems can be valuable scientific aids. They can accelerate code writing and debugging while helping users explore analytical options. However, because their outputs are generated stochastically, they may also produce incorrect, incomplete, or unverified code. As a result, users must still validate, edit, and audit generated outputs, which requires sufficient technical expertise to assess whether the results are correct.
This describes many current widely-available AI systems (e.g., ChatGPT, Claude), which tend to prioritize flexible text and code generation but typically lack domain-specific constraints, standardized workflows, and built-in mechanisms for reproducibility capture. These characteristics limit their suitability for routine execution of scientific analyses and potential for lowering the technical expertise necessary to perform scientific analyses.
Workflow-orchestrating AI: a non-generative alternative for scientific execution
To address these limitations, we propose an alternative architectural pattern for integrating LLMs into scientific workflows, which we term workflow-orchestrating AI. Workflow-orchestrating AI refers to systems in which natural-language interaction is used to route user intent to predefined, domain-specific analytical workflows, rather than to generate new analytical code. In these systems, the LLM functions as an orchestrator, interpreting user requests, selecting appropriate analytical routines, managing inputs, and coordinating execution within a controlled computational environment.
When a user submits a query, the system triggers validated computational routines (e.g., R scripts or workflows written and reviewed by domain experts) rather than producing new code. Execution happens within a controlled runtime and produces structured outputs alongside relevant metadata, such as software and data versions, parameter settings, and execution logs. Importantly, workflow-orchestrating AI is not GenAI since the analytical results are produced by established prewritten scientific software and code, not by generative model output (Figure 1).
Early platforms illustrate how workflow-orchestrating AI can wrap established scientific workflows in a natural-language front end, returning results, explanations, and visualizations while preserving transparent access to the underlying workflow and avoiding reliance on the generation of new code (Figure 1).
This design allows domain experts to embed methodological standards, guardrails, and best practices directly into the system. Thus, results can be reliable, auditable, and reproducible by default, rather than relying on ad hoc code generation followed by manual verification.
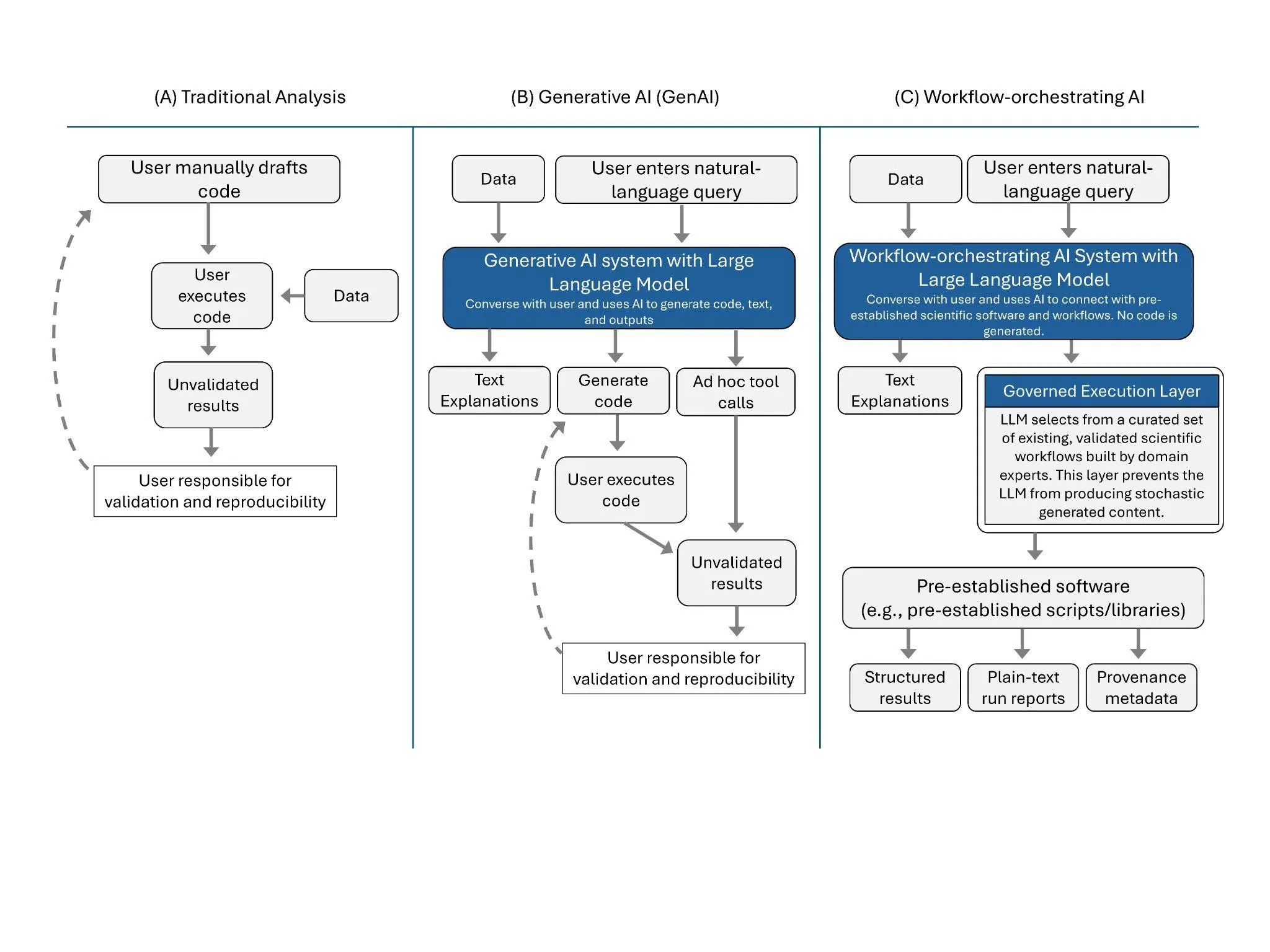
Figure 1. Workflow comparison for three different types of doing ecological analysis: (A) Traditional analysis, which requires users to draft their own code, execute, and troubleshoot it, (B) Generative AI (GenAI), which typically generates ad hoc code using a Large Language Model (LLM) or calls tools in response to a user query, still requiring user knowledge to troubleshoot errors, validate results, and ensure reproducibility, and (C) Workflow-orchestrating AI, which use LLMs to simply select pre-existing scientific software and analytical workflows based on user prompts rather than generating new code.
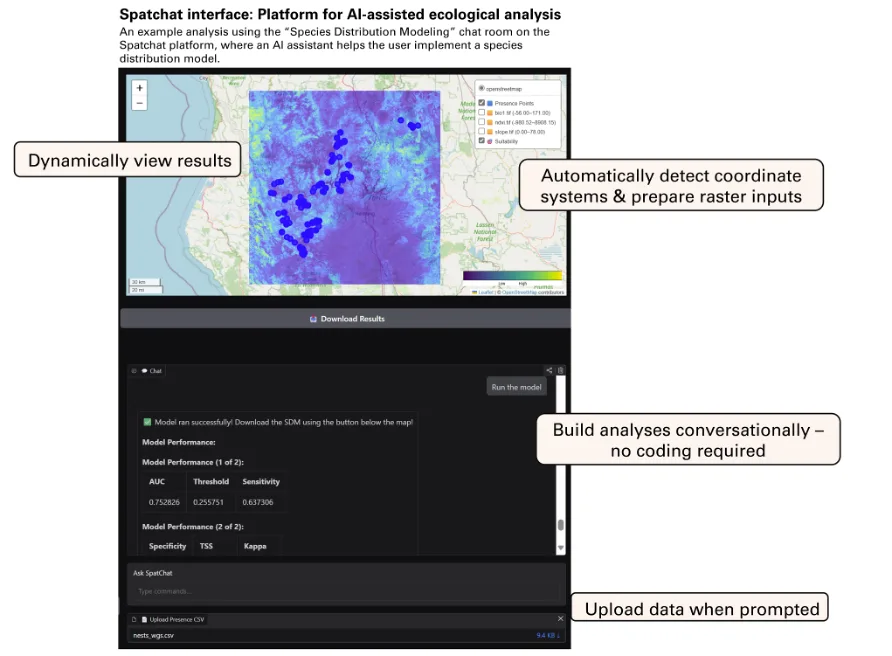
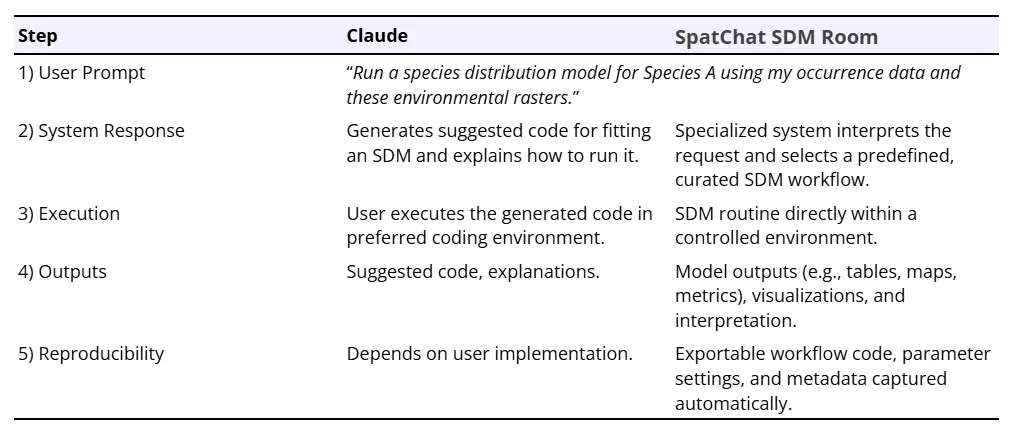
To our knowledge, few ecological platforms currently enable users to execute end-to-end, reproducible spatial analyses entirely through natural language while preserving access to underlying workflows. This excludes tools that primarily generate scripts for users to run independently, as well as fixed graphical interfaces that limit flexibility. SpatChat represents an early example of workflow-orchestrating AI: it wraps established spatial and statistical software in a natural-language front end, constrains execution to validated analytical routines rather than generating new scripts, and preserves exportable outputs, prompts, and code for inspection and reuse. Built as a modular, open-source platform, it allows users to run complex spatial analyses through natural-language interactions that trigger governed analytical routines (Figure 2). SpatChat (www.spatchat.org) is structured using a series of “chat rooms”, each with an AI assistant that specializes in a different analysis. Users can select the chat room relevant to their research question to ask questions, upload data, run analyses, interpret results, and download outputs. For example, users interested in species distributions can explore the species distribution modeling chat room (Figure 2) while users interested in home range analyses can explore the home range analysis chat room. In any given room, a user can type a request into the chatbox and the system will select an appropriate existing workflow to run based on the prompt and data inputs (Table 1). SpatChat is an evolving, community-led project with new features routinely added by contributors. As an early-stage platform, SpatChat does not yet match the robustness or usability of mature analytical software, and some workflows may fail or require refinement; nevertheless, it illustrates the design principles underlying workflow-orchestrating execution systems. The code can be accessed at https://github.com/Spatchat-org.
Lowering barriers and expanding access
The implications of shifting from traditional code-first workflows to governed workflow-orchestrating AI systems are profound. Workflow-orchestrating AI allows researchers to focus on their questions, not code. Researchers can test hypotheses, explore patterns, and run models without spending precious time learning a new package or computer language. By lowering technical hurdles, these tools democratize advanced analysis for students, practitioners, and interdisciplinary collaborators, and deliver sophisticated modeling into classrooms, NGOs, and agencies through intuitive, accessible formats.
Importantly, workflow-orchestrating AI will not replace domain expertise. Instead, they will augment it. Users remain in control of the questions, datasets, and interpretation. Workflow-orchestrating AI simply uses LLMs to select appropriate existing workflows based on user prompts and handle execution in constrained runtimes using existing code written by domain experts, freeing researchers to answer more questions, explore alternatives, communicate results more clearly, and develop stronger foundations in theory.
The ability to engage with large data and modeling through natural languages also opens new opportunities for inclusivity and education. Advanced scientific models can now be introduced to elementary and high school students, allowing them to begin exploring questions they have about advanced scientific topics at a younger age. Community scientists can contribute to spatial prioritization and solve local problems. Small teams without data scientists or GIS specialists can make data-informed decisions. As a result, workflow-orchestrating AI has the potential to broaden who can participate in science and how.
Risks, reproducibility, and the road ahead
As with any transformative technology, the use of workflow-orchestrating AI in science raises valid concerns. Black-box reasoning, version instability, and the risk of ‘hallucinated’ outputs must be acknowledged (Birhane et al. 2023). To mitigate these risks, we emphasize that workflow-orchestrating AI should be LLM-assisted, not LLM-driven. In this context, we use “LLM-assisted” to describe systems in which the language model orchestrates analysis without generating results itself, as in our proposed workflow-orchestrating AI architecture. For example, when a user requests a home range analysis, the LLM itself does not generate code, estimate parameters, or provide outputs; instead it selects an appropriate predefined routine using established scientific libraries (e.g., R packages) within a controlled environment, and reports the results returned by the predefined routine. This separation helps reduce hallucination risk by constraining the LLM’s role to workflow translation and explanation rather than numerical computation and code generation, although validation and user oversight remain essential.
A related category of systems includes AI assistants embedded within existing software, such as spreadsheets or modeling environments, that help users navigate tools through natural language. Workflow-orchestrating AI shares the use of constrained analytical routines with these assistants, but differs in its emphasis on reproducible scientific workflows rather than operating graphical interfaces. In addition, since workflow-orchestrating AI is not specific to any one software, it provides a more generalizable approach to robust LLM-assisted scientific analyses.
Just as importantly, every step must be preserved in underlying code that can be exported, enabling full transparency, reproducibility, and independent verification. Users and reviewers can inspect, export, and rerun workflows with traditional tools. Thus, workflow-orchestrating AI becomes a mechanism for building and communicating robust analyses, not a replacement for scientific integrity.
Reproducibility in workflow-orchestrating AI extends beyond exporting code alone. To support transparent replication and auditing, workflows should also preserve key provenance information, including the analytical routines selected, software and data versions, computation environment details, and relevant LLM configuration parameters (e.g., model version and generation settings). In addition, systems could generate nature language run reports describing the methods selected, specific algorithms invoked, key parameters used, inputs used, and outputs produced. This would enable non-coders to audit decisions and supports replication. Without such documentation, reproducibility risks shifting from traditional code opacity to opaque AI decisions. Designing workflow-orchestrating AI systems that bring methodological choices to the foreground is therefore critical to their scientific legitimacy.
A second, longer-term risk is cognitive offloading. If technical burdens are reduced without parallel attention to training, users may lose familiarity with methodological assumptions, data limitations, and appropriate model choice. Many methodological choices rely on a solid ecological understanding that cannot be delegated to AI. Workflow-orchestrating AI systems should therefore be designed to support learning and critical thinking by making model assumptions clear, offering alternatives, requiring explicit justification for key choices where appropriate, and generating human-readable method summaries alongside results. If designed appropriately, these systems can reduce coding barriers while strengthening methodological literacy.
Importantly, workflow-orchestrating AI will not inherently eliminate costs associated with scientific workflows. In many cases, they will redistribute costs. Reductions in researcher training and implementation time may be offset by sustained engineering effort, infrastructure maintenance, and computational costs associated with deploying and governing LLM-integrated execution systems. The long-term cost-effectiveness of such systems will therefore depend on scale, workflow complexity, model choice, and institutional context. Recognizing these tradeoffs is essential for avoiding over-promising and for aligning system development with realistic scientific needs.
Co-creating a new scientific ecosystem with workflow-orchestrating AI
The contribution of this perspective is to propose a specific design pattern for natural-language AI systems, which we refer to as workflow-orchestrating AI, and propose clear criteria for evaluating such systems. Key criteria include workflow completeness (end-to-end execution), methodological transparency (clear disclosure of model choices and assumptions), reproducibility (code, environment, LLM logs), governance and validation (curated tools and safeguards), usability (how systems behave), and auditability (run reports and structured outputs). Meeting these criteria requires sustained engineering and governance. In practice, the bottleneck might shift from individual researchers writing code to teams maintaining curated workflows, validating updates, managing dependencies, and embedding domain expertise. Successful workflow-orchestrating AI will therefore depend on close collaboration between software engineers and domain scientists with transparent validation processes and community oversight. Comparative benchmarking across platforms is an important step once these systems are fully developed, but beyond the scope of this editorial-style article.
Although we specifically draw on ecological and spatial modeling to illustrate the technical barriers and methodological gap here, many disciplines face similar hurdles. The same solutions presented here can thus be applied and tailored to a diverse set of disciplines. We envision a future where workflow-orchestrating AI is embedded in software, interactive textbooks, and training programs. The goal is not to automate science, but to make scientific tools more accessible, flexible, and responsive while raising the bar for scientists who code to continue developing tools that democratize reproducible science. By embracing AI-driven dialogue and platforms, we can unlock a more inclusive and agile scientific ecosystem.
Acknowledgments
The authors thank Dr. Katie Moriarty, José Juan Rodriguez Gutierrez, Danial Nayeri, and multiple other colleagues for their encouragement and fruitful conversations about the ideas represented in this work.
Author Contributions
Ho Yi Wan: Conceptualization, Writing – review and editing
Logan Hysen: Conceptualization, Writing – original draft
Data Availability
No data was used in this manuscript.
Transparent Peer Review
Results from the Transparent Peer Review can be found here.
Recommended Citation
Wan, H. Y., and L. Hysen. 2026. No coding, no problem: revolutionizing scientific workflows through workflow-orchestrating AI. Stacks Journal: 26005. https://doi.org/10.60102/stacks-26005
References
Birhane, A., A. Kasirzadeh, D. Leslie, and S. Wachter. 2023. Science in the age of large language models. Nature Reviews Physics 5:277–280. https://doi.org/10.1038/s42254-023-00581-4.
Feuerriegel, S., J. Hartmann, C. Janiesch, and P. Zschech. 2024. Generative AI. Business & Information Systems Engineering 66:111–126. https://doi.org/10.1007/s12599-023-00834-7.
Hampton, S. E., C. A. Strasser, J. J. Tewksbury, W. K. Gram, A. E. Budden, A. L. Batcheller, C. S. Duke, and J. H. Porter. 2013. Big data and the future of ecology. Frontiers in Ecology and the Environment 11:156–162. https://doi.org/10.1890/120103.
Moilanen, A., K. A. Wilson, and H. P. Possingham. 2009. Spatial Conservation Prioritization: Quantitative Methods and Computational Tools. Oxford University PressOxford. https://doi.org/10.1093/oso/9780199547760.001.0001.
Ripple, W. J., C. Wolf, T. M. Newsome, P. Barnard, and W. R. Moomaw. 2020. World scientists’ warning of a climate emergency. BioScience 70: 8-12. https://doi.org/10.1093/biosci/biz152.
Sabo, A. N., O. Berger-Tal, D. T. Blumstein, A. L. Greggor, and J. P. Swaddle. 2024. Conservation practitioners’ and researchers’ needs for bridging the knowledge–action gap. Frontiers in Conservation Science 5:1415127. https://doi.org/10.3389/fcosc.2024.1415127.
Dou, D. R., Y. Zhao, J. A. Belk, Y. Zhao, K. M. Casey, D. C. Chen, R. Li, B. Yu, S. Srinivasan, B. T. Abe, K. Kraft, C. Hellström, R. Sjöberg, S. Chang, A. Feng, D. W. Goldman, A. A. Shah, M. Petri, L. S. Chung, D. F. Fiorentino, E. K. Lundberg, A. Wutz, P. J. Utz, and H. Y. Chang. 2024. Xist ribonucleoproteins promote female sex-biased autoimmunity. Cell 187: 733–749. https://doi.org/10.1016/j.cell.2023.12.037.
Soranno, P. A., and D. S. Schimel. 2014. Macrosystems ecology: big data, big ecology. Frontiers in Ecology and the Environment 12:3–3. https://doi.org/10.1890/1540-9295-12.1.3.
Urban, M. C., G. Bocedi, A. P. Hendry, J.-B. Mihoub, G. Pe’er, A. Singer, J. R. Bridle, L. G. Crozier, L. De Meester, W. Godsoe, A. Gonzalez, J. J. Hellmann, R. D. Holt, A. Huth, K. Johst, C. B. Krug, P. W. Leadley, S. C. F. Palmer, J. H. Pantel, A. Schmitz, P. A. Zollner, and J. M. J. Travis. 2016. Improving the forecast for biodiversity under climate change. Science 353:aad8466. https://doi.org/10.1126/science.aad8466.
Open Access
Peer-Reviewed
Creative Commons
Accepted: 02 March 2026
Published: 01 May 2026
No funding was provided for the study.
Conflicts of Interest:
The authors have no conflicts of interest.